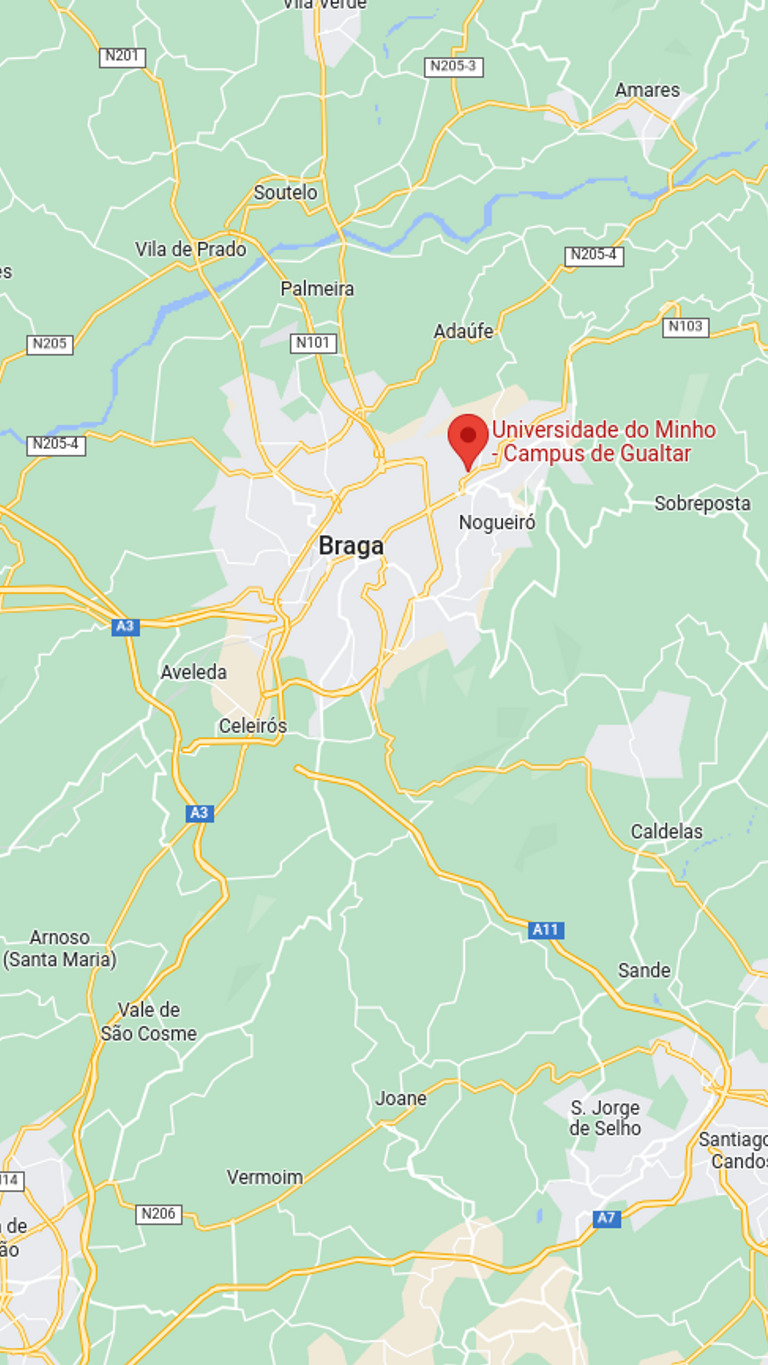
Travel Guide
How to join us
Know before you go! See our travel guide for details on how to get to the event here. Questions? Feel free to ask!
Event Resources
Join us at BOD 2026 for three days of knowledge sharing, innovation and networking in the dynamic field of Bioinformatics.
Registration until March 8
We have options for everyone! Find the best one for you.
NEBiUM Member
17 €Exclusive deal for NEBiUM members.
- Admission for the 3 days of the event
- Informal Dinner (BBQ)
- Workshop registration
Most popular
Student
22,5 €For Bachelor, Master's and PhD students
- Admission for the 3 days of the event
- Informal Dinner (BBQ)
- Workshop registration
Standard
40 €For non-students who want to register for the entire event
- Admission for the 3 days of the event
- Informal Dinner (BBQ)
- Workshop registration
Daily
25 €To register in a particular day
- Single-day admission to all activities (social dinner not included)
- If you choose the 26th:
- Informal Dinner (BBQ)
- If you choose the 27th:
-
Workshop registration
25€ -> 12€
Group Pack
20 € /per personFor a shared experience with others. Available only to parties of 6 or more students
- Includes, for each member:
- Admission for the 3 days of the event
- 10% discount on the registration fee
- Informal Dinner (BBQ)
- Workshop registration
Presenters
75 €For presenters of oral communications, posters and software
- Admission for the 3 days of the event
- Informal Dinner (BBQ)
- Formal Dinner
- Workshop registration
- Presentation at the event